Se = FALSE, color = "black", linetype = 2) +Īnnotate(geom = "text", label = Formula, x = 20, y = 70, size = 6) + 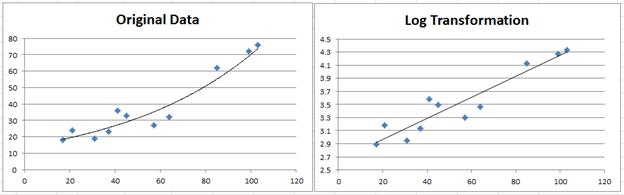
Method.args = list(start = list(a = 8, b = 0.1)), Geom_smooth(aes(group = 1), method = nls, 
Geom_point(stat = "summary", fun = "mean", col = "orange", Ggplot(df, aes(as.numeric(bins) * 10, y)) + 
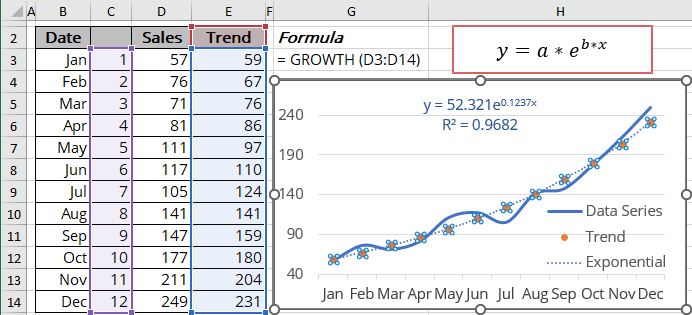
Mod <- nls(y ~ a * exp(b * x), data = df, start = list(a = 8, b = 0.1))įormula <- bquote(y =. The log-transform minimizes the log(Y) residuals whereas the nonlinear regression minimizes the Y residuals.ĭifficult to say without a reproducible example, but your code could be something like this: library(ggplot2) The nls regression does not match the log-transform regression because the two measure the residuals differently. Finally the nls() equation: fit.nls <- nls(Y~a*exp(b*X), start=c(a=5, b=0.2)) The intercept also agrees after we change the scale. Note that the slope value agrees with your Excel example. Now the log transform equation: fit.log <- lm(log(Y)~X) Now we can add a linear regression: fit.linear <- lm(Y~X) First create the data and plot: X <- 1:10 Using the data in the picture, there is no discrepancy between Excel and R.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
